Evolution of Development
One main research focus in the lab is “evo-devo” using arthropods (and their close relatives) as a focal system to investigate the developmental basis of phenotypic change. The serially homologous body plans of arthropods provide an excellent system in which to investigate the origin and diversification of morphological traits, including the origin of novel phenotypes and character identities. I enjoy the challenges that come with establishing new systems, and encourage students to choose their own questions, which often means developing their own organismal system. Thus, in addition to work in the model beetle Tribolium, students in the lab have initiated projects on treehoppers, tardigrades and mayflies.
Testing hypotheses for the origin of morphological novelty in treehoppers
C. R. Fisher, Justin Kratovil, and Elizabeth Jockusch
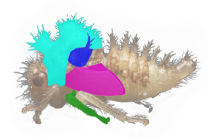 This NSF-funded project investigates the developmental basis for the origin of a fascinating morphological novelty: the highly evolvable pronotal helmet of treehoppers. Evolutionarily, an ancestrally flat part of the dorsal body wall of the wingless first thoracic segment was transformed into a three dimensional cuticular sculpture. This transformation allows treehoppers to take on a wide diversity of shapes, many of which confer mimicry or crypsis. The helmet has been hypothesized to have originated by elaboration of the ancestral body wall developmental patterning network or alternatively by cooption of the wing or leg patterning network. These hypotheses make specific predictions about patterns of similarity and difference in gene expression and function across tissue types and species. This project combines comparative transcriptomics with the analysis of gene function in an evolutionary context to test these predictions.
This NSF-funded project investigates the developmental basis for the origin of a fascinating morphological novelty: the highly evolvable pronotal helmet of treehoppers. Evolutionarily, an ancestrally flat part of the dorsal body wall of the wingless first thoracic segment was transformed into a three dimensional cuticular sculpture. This transformation allows treehoppers to take on a wide diversity of shapes, many of which confer mimicry or crypsis. The helmet has been hypothesized to have originated by elaboration of the ancestral body wall developmental patterning network or alternatively by cooption of the wing or leg patterning network. These hypotheses make specific predictions about patterns of similarity and difference in gene expression and function across tissue types and species. This project combines comparative transcriptomics with the analysis of gene function in an evolutionary context to test these predictions.
Diversification of insect appendages and the evolution of serial homologues
Elizabeth Jockusch, with Frank Smith and Dave Angelini
 Arthropod appendages diversified from an ancestrally homonomous series, assuming distinct, conserved identities early in arthropod evolution. Using our main arthropod lab model, the flour beetle Tribolium castaneum, we have analyzed conservation vs. divergence in appendage patterning at multiple levels: across developmental stages within species (i.e., embryonic vs. metamorphic patterning), across appendage types within species (e.g., mouthparts vs. legs), and across directly homologous appendage types in different species (e.g., by comparing Tribolium to Drosophila). Results of this work include evidence for “concerted evolution” of patterning across divergent appendage types, a model for patterning of ancestral mouthpart morphologies, and evidence for divergence in how appendage identities are developmentally regulated both across species and across developmental stages.
Arthropod appendages diversified from an ancestrally homonomous series, assuming distinct, conserved identities early in arthropod evolution. Using our main arthropod lab model, the flour beetle Tribolium castaneum, we have analyzed conservation vs. divergence in appendage patterning at multiple levels: across developmental stages within species (i.e., embryonic vs. metamorphic patterning), across appendage types within species (e.g., mouthparts vs. legs), and across directly homologous appendage types in different species (e.g., by comparing Tribolium to Drosophila). Results of this work include evidence for “concerted evolution” of patterning across divergent appendage types, a model for patterning of ancestral mouthpart morphologies, and evidence for divergence in how appendage identities are developmentally regulated both across species and across developmental stages.
Diversification and Speciation
Another main research focus in the lab is the process of lineage diversification. We combine molecular phylogenetics with analyses of morphological variation, and occasionally also field nad lab experiments. My interest in molecular phylogenetics grew out of my dissertation work on phenotypic plasticity in slender salamanders, and the need for a phylogeny on which to map the reaction norm variation I discovered. Suffice it to say the evolutionary history of the group turned out to be much more complex (and interesting) than anticipated!
Speciation and reticulation in California herps
Elizabeth Jockusch and Nick Van Gilder
collaborators: Jonathan Richmond (US Geological Survey), Chris Norment (Suny Brockport), Steve Parmenter (Cal Fish and Wildlife), Robert Hansen, Sam Sweet, and Iñigo Martínez-Solano (National Museum of Spain)
 The complex climatic and geological history of western North America has produced numerous opportunities for both isolation and secondary contact across an extended evolutionary timeframe. We use two lineages endemic to western North America as natural experiments for testing hypotheses about diversification. Currently, we are particularly interested in the role of reticulation in evolution of these lineages. The slender salamanders, genus Batrachoseps, are the most diverse salamander clade in western North America, and show extreme patterns of microendemism across much of their range. They also display interesting patterns of convergent and divergent morphological evolution. Skinks of the Plestiodon skiltonianus complex provide a hypothesized example of parallel speciation, an unusual speciation mode in which the same reproductive isolating mechanism originates multiple times in parallel, allowing for reticulation. Both species complexes display in situ diversification, persistence of ancient lineages, and evidence for multiple reticulation events. Our aim is to infer the evolutionary history of these lineages from geographically comprehensive sets of samples, which will allow us to delimit evolutionary units; identify contact zones and characterize gene flow between well-differentiated taxa; and evaluate the extent to which phylogenetic trees vs. phylogenetic networks (resulting from the addition of reticulation to phylogenetic trees) describe the diversification process. Andrew is also studying why tail color has diverged between skink populations.
The complex climatic and geological history of western North America has produced numerous opportunities for both isolation and secondary contact across an extended evolutionary timeframe. We use two lineages endemic to western North America as natural experiments for testing hypotheses about diversification. Currently, we are particularly interested in the role of reticulation in evolution of these lineages. The slender salamanders, genus Batrachoseps, are the most diverse salamander clade in western North America, and show extreme patterns of microendemism across much of their range. They also display interesting patterns of convergent and divergent morphological evolution. Skinks of the Plestiodon skiltonianus complex provide a hypothesized example of parallel speciation, an unusual speciation mode in which the same reproductive isolating mechanism originates multiple times in parallel, allowing for reticulation. Both species complexes display in situ diversification, persistence of ancient lineages, and evidence for multiple reticulation events. Our aim is to infer the evolutionary history of these lineages from geographically comprehensive sets of samples, which will allow us to delimit evolutionary units; identify contact zones and characterize gene flow between well-differentiated taxa; and evaluate the extent to which phylogenetic trees vs. phylogenetic networks (resulting from the addition of reticulation to phylogenetic trees) describe the diversification process. Andrew is also studying why tail color has diverged between skink populations.
Diversification of parasites and their hosts
Janine Caira, Hannah Ralicki, Jill Wegrzyn and Elizabeth Jockusch
collaborators: Kirsten Jensen, Mark Holder and Jeet Sukumaran (University of Kansas)
Parasites are ubiquitous in nature, and often show tight associations with their hosts. A common approach to understanding the evolution of these associations is to assume that parasite diversification tracks host diversification, with other explanations (e.g., events such as host switching) invoked only secondarily to explain mismatches. However, even in taxa that show strong host specificity, matched patterns of phylogenetic divergence often do not occur. This NSF-funded project aims to develop a new framework for testing the multiple biotic and abiotic factors that structure the phylogenetic relationships of parasites and their hosts. We use trophically-transmitted tapeworms that inhabit the spiral intestines of elasmobranchs as our empirical system, and are analyzing parasite clades that display different degrees of cophylogenetic structure. My lab collaborates on the molecular data collection and phylogenetic analyses, including contributing to the development of genomic resources and novel markers for cestode phylogenetics.
The ecological and evolutionary significance of color morphs in Plethodon cinereus
Annette Evans, Elizabeth Jockusch and Mark Urban
 Red-backed salamanders (Plethodon cinereus) display two color morphs across much of their range in eastern North America. Geographic variation in the relative frequency of these color morphs, combined with their long-term persistence, suggests that some form of balancing selection maintains the morphs. Extensive prior work on the system has led to many hypotheses about the possible fitness differences correlated with/linked to the different phenotypes, but no hypothesis fully accounts for the observed spatial patterns. Annette’s dissertation research incorporates resurvey work to test whether morph frequencies in New England have evolved in response to climate or land use change, along with tests for physiological differences between the morphs and for plasticity in morph development in response to embryonic temperature.
Red-backed salamanders (Plethodon cinereus) display two color morphs across much of their range in eastern North America. Geographic variation in the relative frequency of these color morphs, combined with their long-term persistence, suggests that some form of balancing selection maintains the morphs. Extensive prior work on the system has led to many hypotheses about the possible fitness differences correlated with/linked to the different phenotypes, but no hypothesis fully accounts for the observed spatial patterns. Annette’s dissertation research incorporates resurvey work to test whether morph frequencies in New England have evolved in response to climate or land use change, along with tests for physiological differences between the morphs and for plasticity in morph development in response to embryonic temperature.
Other Lab Projects
I am open to sponsoring students or postdocs working on any question about phenotypic evolution for which I have relevant expertise. Below is a brief overview of past projects initiated in the lab.
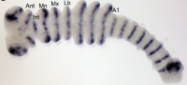 Ph.D. student Brigid O’Donnell combined molecular phylogenetics with analyses of morphological variation to investigate patterns of evolution in mayfly gills, focusing on the family Leptophlebiidae. She also established mayflies as a developmentally tractable system, obtaining embryos from mass emergences of the species Ephoron leukon, rearing them through to hatching in the lab, and using components of the segmentation gene network to develop methods for characterizing gene expression.
Ph.D. student Brigid O’Donnell combined molecular phylogenetics with analyses of morphological variation to investigate patterns of evolution in mayfly gills, focusing on the family Leptophlebiidae. She also established mayflies as a developmentally tractable system, obtaining embryos from mass emergences of the species Ephoron leukon, rearing them through to hatching in the lab, and using components of the segmentation gene network to develop methods for characterizing gene expression.

Ph.D. student Roberta Engel studied diversification of pseudoscorpions in the genus Synsphyronus on granite outcrops that form a terrestrial archipelago embedded in an agricultural landscape in southwestern Australia. Over three field seasons in Australia, hosted by Mark Harvey’s lab, she sampled more than 100 outcrops, as well as pseudoscorpions from throughout the country. She discovered both high morphological and genetic diversity in the outcrop region. Phylogenetic analyses showed that the outcrops were colonized multiple times, with rock-restricted lineages undergoing subsequent diversification across outcrops.
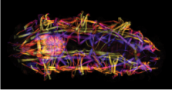 Ph.D. student Frank Smith combined analyses of metamorphic appendage patterning and identity determination in Tribolium with a new project using tardigrades to test hypotheses about the origin of arthropod body plans. Tardigrades share with arthropods a segmented body plan with one pair of appendages per segment, but retain key ancestral traits including homonomous unjointed appendages. Frank solved the long-standing challenge of developing a robust in situ hybridization protocol for tardigrades, and used his method to characterize Distal-less expression in Hypsibius dujardini. He also characterized muscle and nervous system morphology in this species, allowing him to uniquely identify each body segment.
Ph.D. student Frank Smith combined analyses of metamorphic appendage patterning and identity determination in Tribolium with a new project using tardigrades to test hypotheses about the origin of arthropod body plans. Tardigrades share with arthropods a segmented body plan with one pair of appendages per segment, but retain key ancestral traits including homonomous unjointed appendages. Frank solved the long-standing challenge of developing a robust in situ hybridization protocol for tardigrades, and used his method to characterize Distal-less expression in Hypsibius dujardini. He also characterized muscle and nervous system morphology in this species, allowing him to uniquely identify each body segment.
Postdoc Dave Angelini initiated our RNAi-screen of candidate gene functions during appendage patterning in Tribolium. His initial focus was on antennal patterning, since antennal morphology varies greatly across species. This work identified a suite of candidate genes whose functions differ between Drosophila and Tribolium, and that may play roles in the evolution of antennal morphology. He also expanded our work to include multiple species of Tribolium and produced a phylogeny for these species.